easyMF: A Web Platform for Matrix Factorization-Based Gene Discovery from Large-scale Transcriptome Data
Abstract
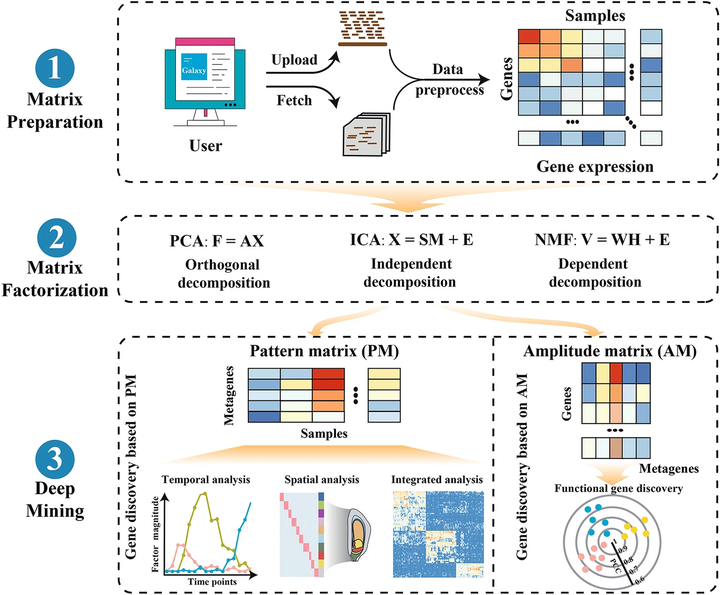
With the development of high-throughput experimental technologies, large-scale RNA sequencing (RNA-Seq) data have been and continue to be produced, but have led to challenges in extracting relevant biological knowledge hidden in the produced high-dimensional gene expression matrices. Here, we develop easyMF (https://github.com/cma2015/easyMF), a web platform that can facilitate functional gene discovery from large-scale transcriptome data using matrix factorization (MF) algorithms. Compared with existing MF-based software packages, easyMF exhibits several promising features, such as greater functionality, flexibility and ease of use. The easyMF platform is equipped using the Big-Data-supported Galaxy system with user-friendly graphic user interfaces, allowing users with little programming experience to streamline transcriptome analysis from raw reads to gene expression, carry out multiple-scenario MF analysis, and perform multiple-way MF-based gene discovery. easyMF is also powered with the advanced packing technology to enhance ease of use under different operating systems and computational environments. We illustrated the application of easyMF for seed gene discovery from temporal, spatial, and integrated RNA-Seq datasets of maize (Zea mays L.), resulting in the identification of 3,167 seed stage-specific, 1,849 seed compartment-specific, and 774 seed-specific genes, respectively. The present results also indicated that easyMF can prioritize seed-related genes with superior prediction performance over the state-of-art network-based gene prioritization system MaizeNet. As a modular, containerized and open-source platform, easyMF can be further customized to satisfy users’ specific demands of functional gene discovery and deployed as a web service for broad applications.