Welcome to deepTFBS !
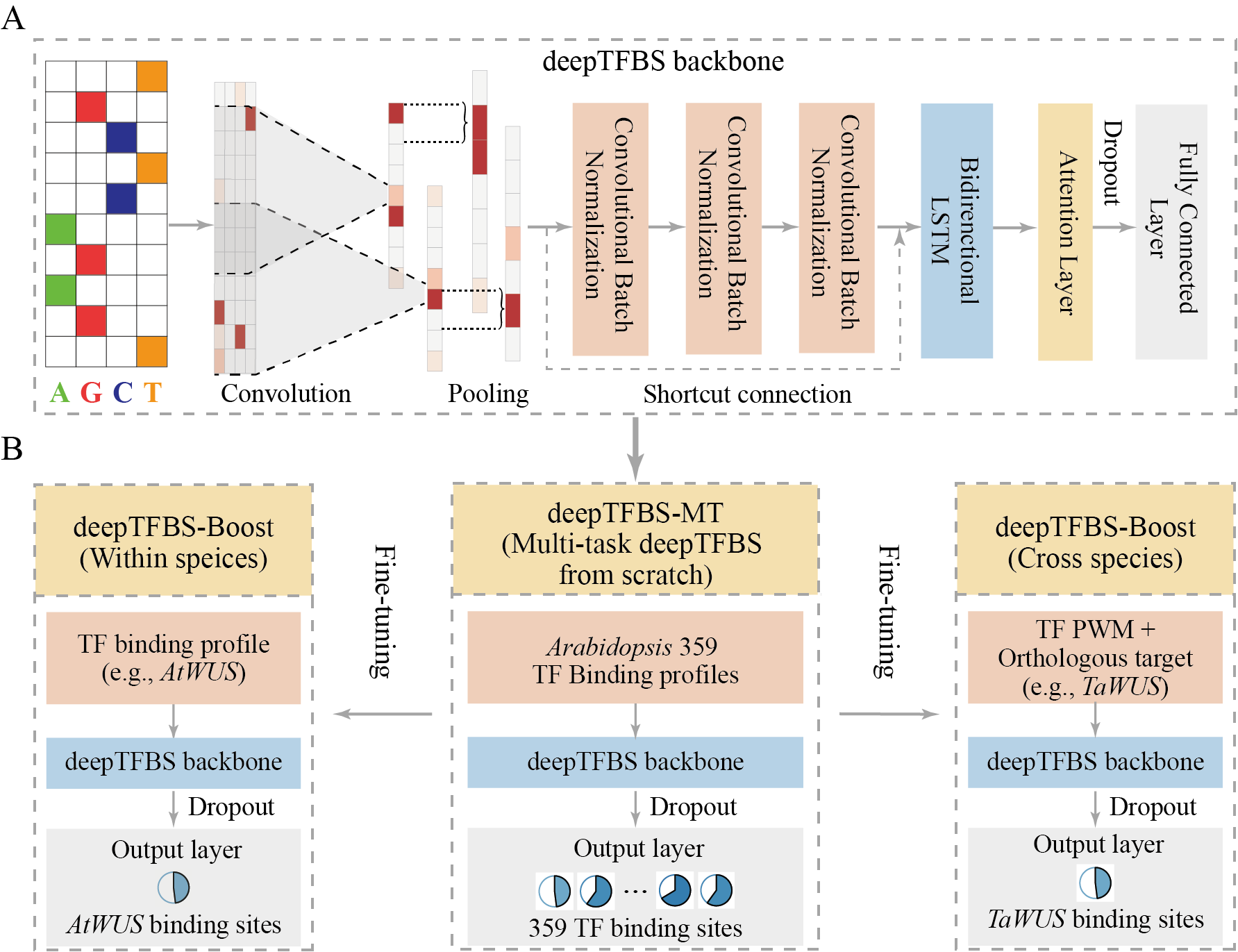
Precise prediction of transcription factor binding sites (TFBS) is crucial for understanding gene regulation mechanisms. Here, we introduce deepTFBS, a comprehensive deep learning framework that builds robust DNA language models of TF binding grammars for accurately predicting TFBS both within and across plant species. deepTFBS takes advantages of multi-task learning technique to integrate large-scale TF binding profiles for pre-training, and is capable of leveraging knowledge from pre-trained models via transfer learning, representing an innovation in that it can improve prediction accuracy of TFBS under small-sample training and cross-species prediction tasks. When tested on the binding data of 359 Arabidopsis TFs, deepTFBS outperformed deepSEA for 323 TFs and surpassed DanQ for 246 TFs, as measured by the area under the precision-recall curve (PRAUC). Furthermore, deepTFBS has the capability of utilizing information from gene conservation and binding motifs, providing an efficient way of TFBS prediction for species lacking sequencing binding data. A case study focusing on the WUSCHEL (WUS) transcription factor further illustrates deepTFBS’s cross-species application potential from Arabidopsis to wheat. Through yeast one-hybrid assays, seven of fourteen randomly slected predicted WUS targets in wheat were experimentally validated. deepTFBS is provided in open source (https://github.com/cma2015/deepTFBS) with a web server and Docker image to support the dissection of gene regulation in plants and other organisms.